Database Performance for Genomic Data at Scale
Many genomic databases applications still use relational models, but NoSQL databases can increase genomic data storage efficiency.

Genomic data growth continues to accelerate and the use of these for research and clinical purposes is constantly expanding. Initially, the use of genomic sequencing in the clinical setting was focused on human genetics to identify risks for and causes of human disease. As sequencing costs declined, the use of genomics has expanded beyond the human genome and is now also used to identify pathogens responsible for infection and, with the COVID-19 pandemic, used to surveil for new viral strains and identify mutations that may lead to increased transmissibility, pathogenicity, or treatment resistance.
With the continued growth in genomic data and annotations comes the need to develop approaches to efficiently store, manage, and access these data. While there is rarely a one size fits all option, exploring different technologies for specific use cases can reduce costs and increase responsiveness in end-user applications.
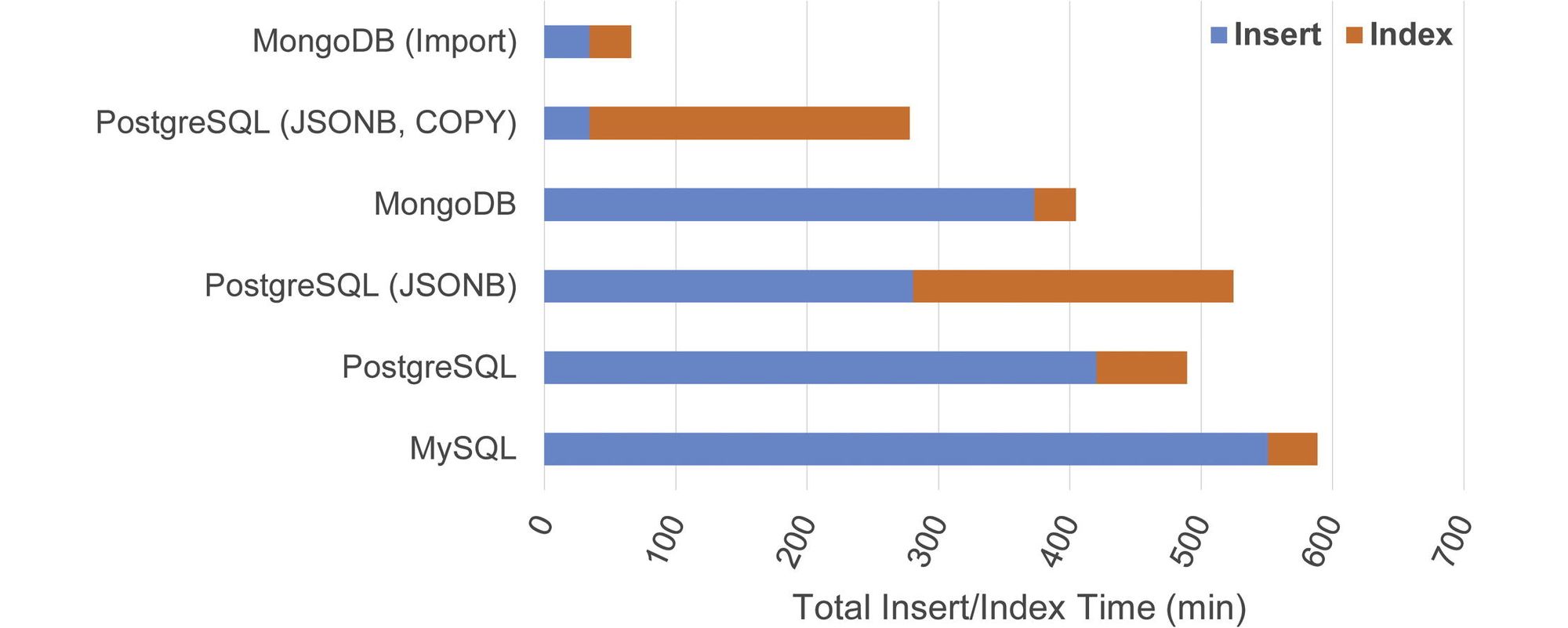
Many applications still default to relational database models because of their familiarity. But from experiments we performed several years ago (https://github.com/wadeschulz/research_snpdb) demonstrated that NoSQL databases, such as MongoDB, can reduce storage and access times for genomic annotation data (https://www.sciencedirect.com/science/article/pii/S1532046416301526).
Your software architecture, data model, and hardware can all affect the performance of data storage and access. But for high volume applications, studying the performance of underlying technology choices is critical, as implementing the wrong solution early in the development course can be much more costly to fix in the future.